Introduction
Electrocardiogram (ECG) is a visual time series of cardiac activity and is an important basis for diagnosing heart disease.
Traditional ECG interpretation relies on manual analysis. Clinicians use accumulated experience and domain knowledge to produce accurate diagnoses.
With a high incidence of cardiac disease and large daily volumes of ECGs, manual diagnosis is inefficient and can lead to misdiagnosis, negatively affecting patients and healthcare providers. Automatic ECG classification methods have therefore been developed to address these issues.
This work designs a hardware ECG data acquisition system based on an STM32 microcontroller for signal acquisition and transmission. The BMD101 chip is used to acquire raw ECG signals, and data are transmitted to a PC application and an Android app via Bluetooth and WiFi for classification processing.

1. Current Research on ECG Classification Methods
There are two main approaches to ECG classification: waveform-shape based methods and feature-based methods.
Waveform-shape based methods extract characteristic waveforms and apply medical classification rules. Philip Deehazal et al. used 12 morphological and interval features to train and test a classifier, dividing heartbeats into five categories with an accuracy of 0.81. Li Kunyang et al. combined wavelet transform and mathematical morphology to detect QRS feature points, obtaining QRS width and RR interval parameters; combined with clinical experience, they classified beats into four categories with an accuracy of 0.94. Shape-based methods require high signal quality and have limited noise robustness; they also require precise detection of waveform feature points and are not well suited for dynamic ECGs. As a result, the extracted feature vectors are often insufficient, limiting the number of diagnosable categories and lowering accuracy.
Feature-based methods are the most widely used. Liu Shixiong used wavelet analysis to locate QRS complexes, extracted 26 features from each QRS complex, and applied fuzzy clustering for classification. Luo Dehan et al. applied a multi-order feedforward artificial neural network to achieve six-class ECG classification.
Ji Hu extracted features using multiple discriminants and principal component analysis, then used support vector machines for classification. Yusn et al. used independent component analysis in the time domain to extract ECG features and applied neural networks for six-class classification with good results.
Osowski designed a cascade classifier combining a fuzzy self-organizing layer and a multilayer perceptron to achieve seven-class ECG classification with 96% accuracy. Owis analyzed ECGs in the Fourier transform domain, extracted feature vectors, and used a nearest-neighbor classifier; experiments reported 100% recognition for five cardiac condition types, while recognition for other conditions was lower.
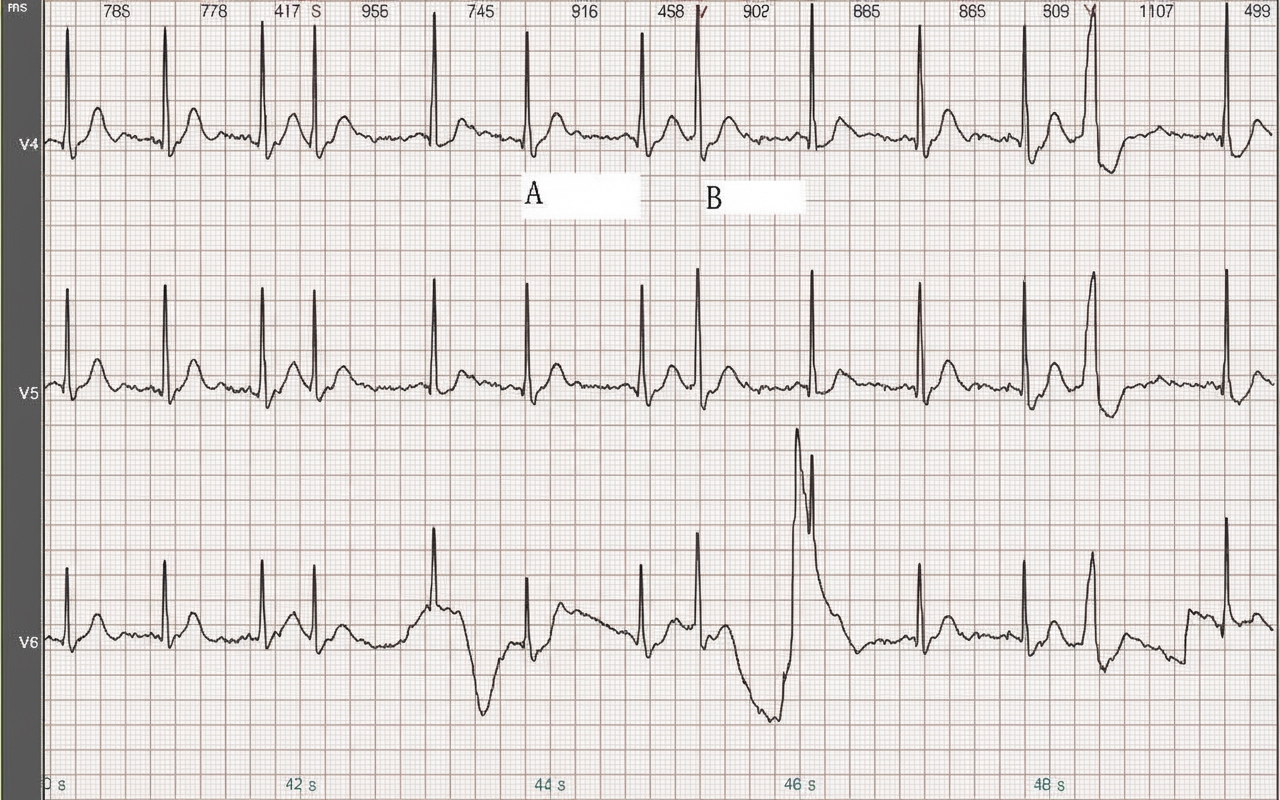
2. Hardware Circuit Design
2.1 System Overview
The ECG monitoring system comprises four modules: ECG data acquisition, wireless transmission, Android app for data reception and processing, and a PC application for data reception and processing.
When a user experiences cardiac discomfort, the device can immediately collect ECG signals. Collected data are transmitted via Bluetooth to an Android app, which can display ECG waveforms and heart rate in real time, determine the user's geographic location, and upload data to a server.
2.2 Acquisition Module
ECG Acquisition Chip
The BMD101 chip integrates an advanced analog front end and a flexible digital signal processing architecture. It can acquire bio-signals from microvolts to millivolts and process them with Neurosky's proprietary algorithms. At its core is a system management unit responsible for overall system configuration, runtime management, internal and external communication, proprietary algorithm computation, and power management. BMD101 also includes an embedded DSP to accelerate various digital filtering computations under the system management unit's control.
3. Classification Algorithms and System Software Design
This work uses the open-source deep learning framework Faster R-CNN for object detection. Training specific network models under this framework requires GPU resources. For ZF net and VGG-16 net models, GPU memory requirements are at least 3 GB and 8 GB, respectively.
3.1 Dataset Construction
Dataset construction includes building the training and test sets. Training set construction involves extracting 44 sets of human ECG data, R-peak detection, heartbeat image cropping, beat labeling, and data augmentation via translation and rotation. The augmented heartbeat image set is used as the training set. The test set is constructed by collecting ECG data, preprocessing, R-peak detection, and cropping heartbeat images to create an unlabeled test set.
3.1.1 R-peak Detection
Wavelet transform techniques perform well in improving Q-wave detection accuracy. According to wavelet transform theory, the R-peak corresponds to a zero crossing between a local modulus maximum and minimum with a stable displacement relationship. Using wavelet transform to detect QRS complexes locates R waves by identifying maxima of the modulus; smaller scales are then searched forward and backward from the R peak to determine QRS start and end points, avoiding width distortion. For noisy or atypical R waves, one must refine modulus maxima points to reduce noise interference.
In this experiment, discrete wavelet transform is used to detect R waves. First, modulus maxima are detected, then zero crossings are found to identify QRS locations. An adaptive noise threshold is introduced to determine whether a detected spike is an R wave or an artifact. or each R-peak detected by wavelet transform, record the R coordinate, search 250 ms to the left from the R point; if a valid point exists, record the left endpoint, otherwise stop. Then search 250 ms to the right of the R point; if a valid point exists, record the right endpoint, otherwise stop. Crop the waveform between these endpoints to create the sample input for the deep learning network.
3.2 Training the Classification Model
In supervised deep learning, labeled datasets are fed to a network according to a strategy. Over thousands of iterations, the model adjusts internal parameters to produce an effective classifier.
In this work, the training set is stored as XML files. Each image is an independent variable xi; the bounding box coordinates [x1,y1,x2,y2] and the class label Label (Label ∈ {“N”, “S”, “V”, “F”}) are dependent variables yi. Analogous to fitting a line to many points in two-dimensional space, many (xi, yi) pairs in deep learning fit a complex function determined by model parameters.
Thus, the goal is to train a deep network to represent a complex nonlinear function with many parameters to predict labels given inputs.
Within the Faster R-CNN framework, several parts must be modified for training:
- Update dataset paths, labels, and other hyperparameter files.
- Adjust the number of classes in train_val.prototxt and test.prototxt.
- Modify the solver file and set its path.
- Run the training script script_faster_rcnn_VOC2007_ZF.m to execute training.
- After training, modify the generated detection_test.prototxt file.
For parameter settings, default values are used except for max_iter and stepsize in the solver file. max_iter is the maximum number of training iterations: too small causes undertraining; too large may cause overfitting. stepsize is the iteration count after which the learning rate decays. The learning rate is critical: a larger rate early in training speeds convergence; a smaller rate later prevents overshooting the optimum.
The dataset used contains 800 images. Multiple training experiments found best results with max_iter = 4000 and stepsize = 3000.
4. Classification Performance Testing
The Faster R-CNN framework produces three outputs for heartbeat classification: bounding box (box), class label, and score. The box is defined by coordinates of the top-left and bottom-right corners, represented as a 1×4 vector. The class label is the model's class prediction for that box and, in this work, takes one of four discrete values. .
5. System Integration and Conclusion
A hardware ECG acquisition system based on an STM32 microcontroller was designed for signal acquisition and transmission. The BMD101 chip acquires raw ECG signals, and Bluetooth and WiFi transmit data to a PC application and an Android app. A PC application was implemented for ECG display and storage, and an Android app implements real-time ECG display, user location, and remote server upload of ECG data.
Accurate ECG classification depends on extracting correct features. This work trains a deep neural network directly on preprocessed ECG signals, extracting features layer by layer and fitting an automatic ECG classification model. The trained model was used to classify ECGs and produced satisfactory results.